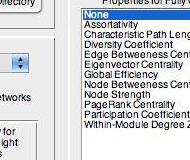
This Matlab toolbox runs a GLM on graph theoretic network properties in brain networks. The GLM accepts continuous & categorical between-participant predictors and categorical within-participant predictors. Significance is determined via non-parametric permutation tests. The toolbox allows testing of both fully connected and thresholded networks (based on a range of thresholds).
The toolbox also provides a data processing path for resting state fMRI data. Several options for partialing nuisance signals are available, including local and total white matter signal (Jo et al., 2013), calculation of Saad et al. (2013)'s GCOR, and the use of Chen et al. (2012) GNI method to determine whether global signal partialing is needed. In addition, Power et al. (2014)'s motion scrubbing method is available.
Features
- GLM for graph theoretic properties
- Processing stream for resting state fMRI