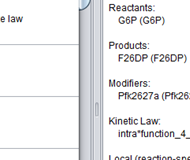
The SBML Reaction Finder retrieves and extracts individual chemical reactions from curated SBML models.
Biological modelers who want to build chemical network models using components from a repository of existing, published reactions can use this tool to access the thousands of individual reactions in the BioModels repository.
The software is open source, and is part of the SBP group's modular, semantics-based approach to biosimulation modeling. Dr. Herbert Sauro at the University of Washington's Bioengineering Department guided its development.
The SBML Reaction Finder knowledge base To retrieve individual reactions of interest, the SRF searches a custom knowledge base, coded in the Web Ontology Language (OWL), that stores the biological annotations, unique identifiers, and source model affiliation for each reaction in BioModels. Although RDF would be expressive enough to capture the information in our knowledge base, we chose to use OWL so that, as the tool grows in sophistication, we will have the opportunity to explore automated reasoning as a means of improving model retrieval and extraction. At the time of manuscript preparation, the SBML Reaction Finder’s knowledge base contained a total of 6,533 reactions, 5,648 (86%) of which are associated with a GO term, either through direct annotation against GO or through Enzyme Commission number cross-references. Each time the user performs a search with the SBML Reaction Finder, the program looks for pattern matches in the names and IDs of the reactions in this knowledge base, the preferred names and synonyms of any GO annotations, the names of their parent models, and the names and synonyms of the taxon in which they occur. Users can update this knowledge base at any time to ensure that they are searching over the most up-to date version of the BioModels database. The SBML Reaction Finder update tool makes use of BioModels, UniProt, BioPortal and BRENDA web services to collect all the curated SBML models in the repository, classify their reactions according to their GO annotations, and store the human-readable names of these annotations for easier retrieval of individual reactions.
Easily find and extract specific chemical reactions from the BioModels database.
Features of SBML Reaction Finder:
- Autosuggest for Gene Ontology terms
- Update feature ensures access to the latest version of BioModels repository